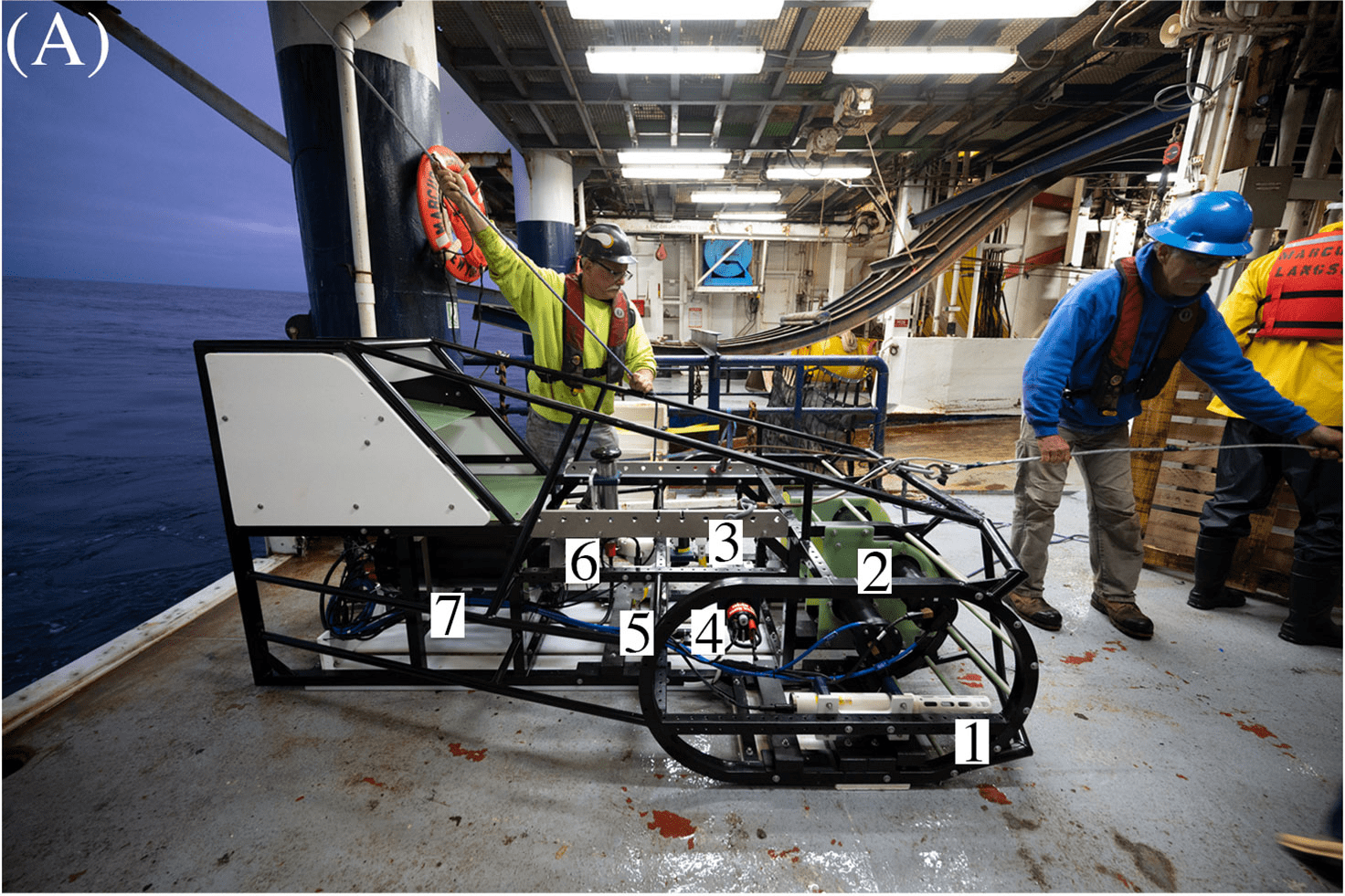
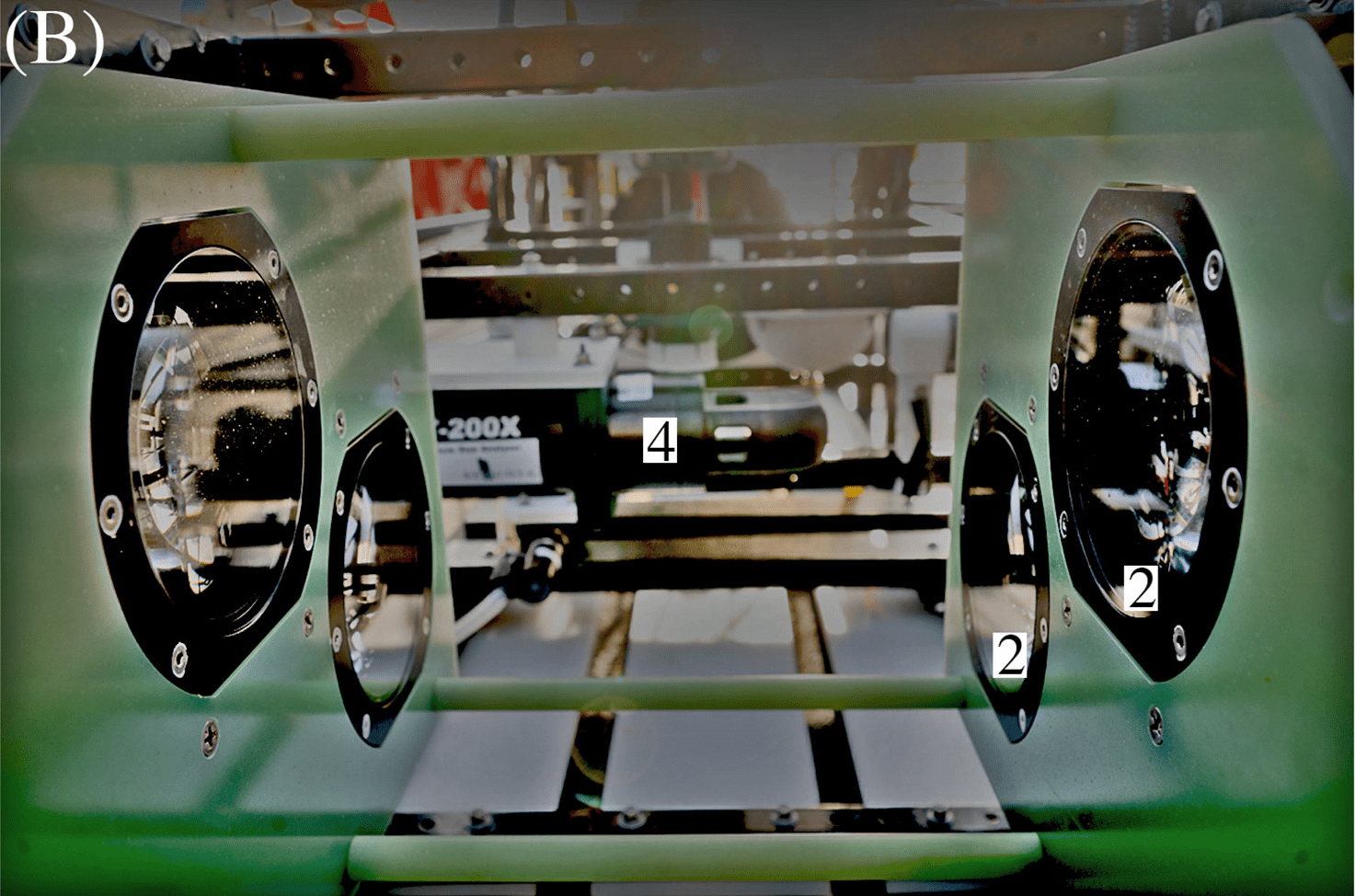
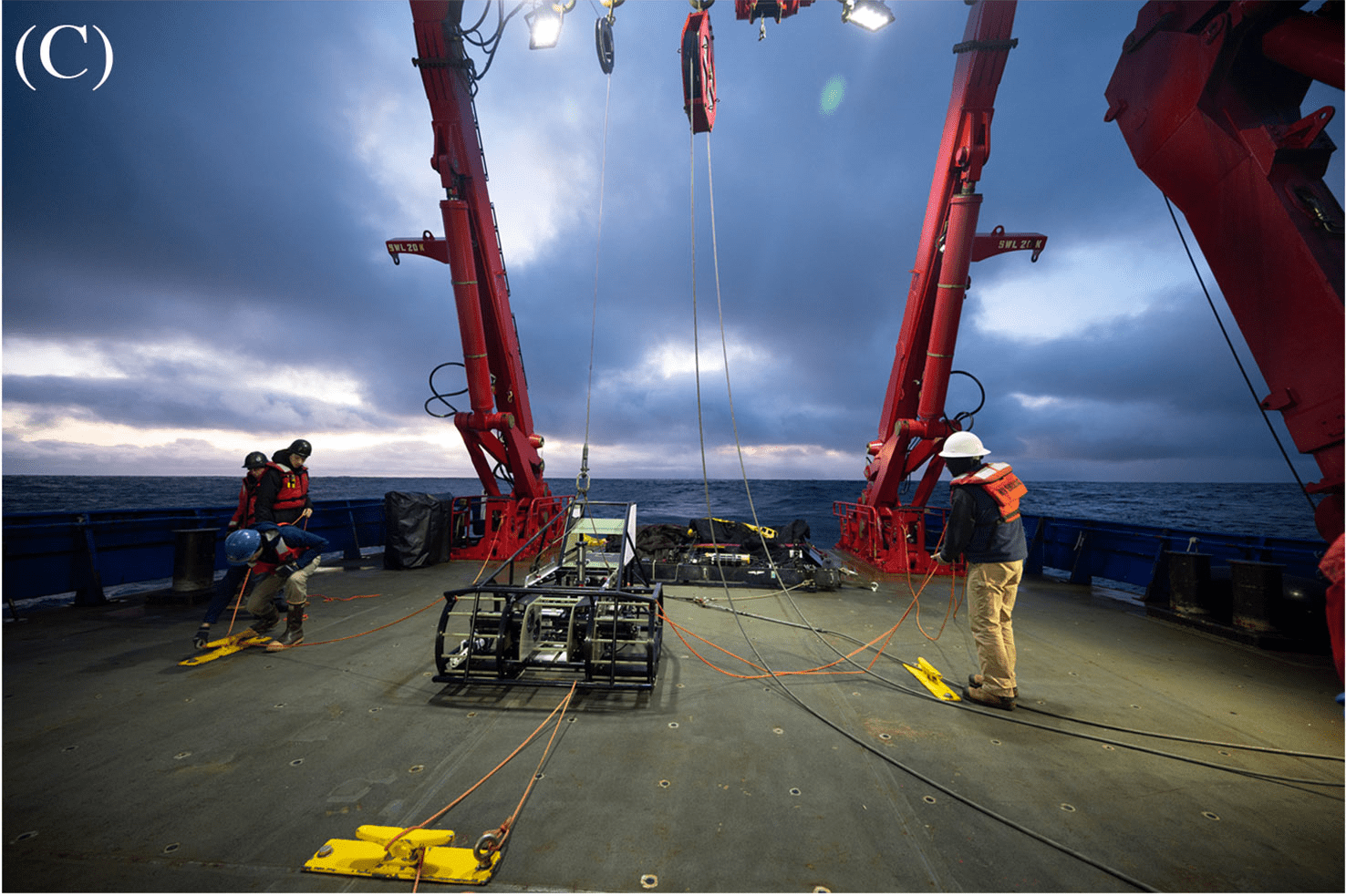



ISIIS-3 and its components (A) Lateral view; (B) Close-up of the two stacked Bellamare ISIIS-DPI-125 camera units; (C) Deployments of the detector from the sides of the ship, using a crane. Photo credit: Ellie Lafferty.
Vital Base of Ocean Food Web Can Now Be Studied in Bulk Using AI
The ocean provides about half of the oxygen necessary for humans to survive. Scientists at Oregon State University wanted to better understand how plankton populations in the ocean will respond to climate change. To study plankton populations, they developed a new way of photographing large populations of these creatures at sea, using PSC’s Bridges-2 system to analyze and identify plankton. They’ve now adapted this AI classifier to use in an onboard computer, giving them the ability to redirect their cruises on the fly, with deeper analysis using Bridges-2 back on land.
WHY IT’S IMPORTANT
To a large extent, we survive by the ocean’s bounty. The world’s fisheries produce 17% of humanity’s per capita protein intake. More importantly, half of our oxygen comes from plants and plankton in the seas. That’s why scientists are trying to understand plankton, the foundation of that food and oxygen production, better. Plankton includes all living things that float in the ocean with limited ability to swim. In addition to tiny plants, or phytoplankton, they include zooplankton, animals like jellyfish and the larvae of fish too small to propel themselves any large distance — as small as 0.2 mm (a little less than one one-hundredth of an inch). Microscopic plant life like algae get eaten by zooplankton; they in turn feed smaller fish, who are eaten by larger fish. Baleen whales skip the line by feeding on zooplankton directly — but in either case, all those animals would not be there without plankton starting the chain.“We know a lot about [the ocean food chain], about how it’s structured … We don’t know in particular how responsive … the food chain will be … to situations we are encountering now — heat waves, warmer waters, acidification, low oxygen — driven by climate change.” — Robert Cowen, Oregon State University
What scientists don’t understand is how this food chain works in three dimensions. The traditional way of sampling plankton — basically trailing a net behind your ship — mixes up species from different depths, so you can’t be sure that the predator and prey you’ve caught live in the same place, or just happened to be scooped together. Worse, we don’t know how that complex system will change, or if it could even collapse, as the oceans warm and acidify.
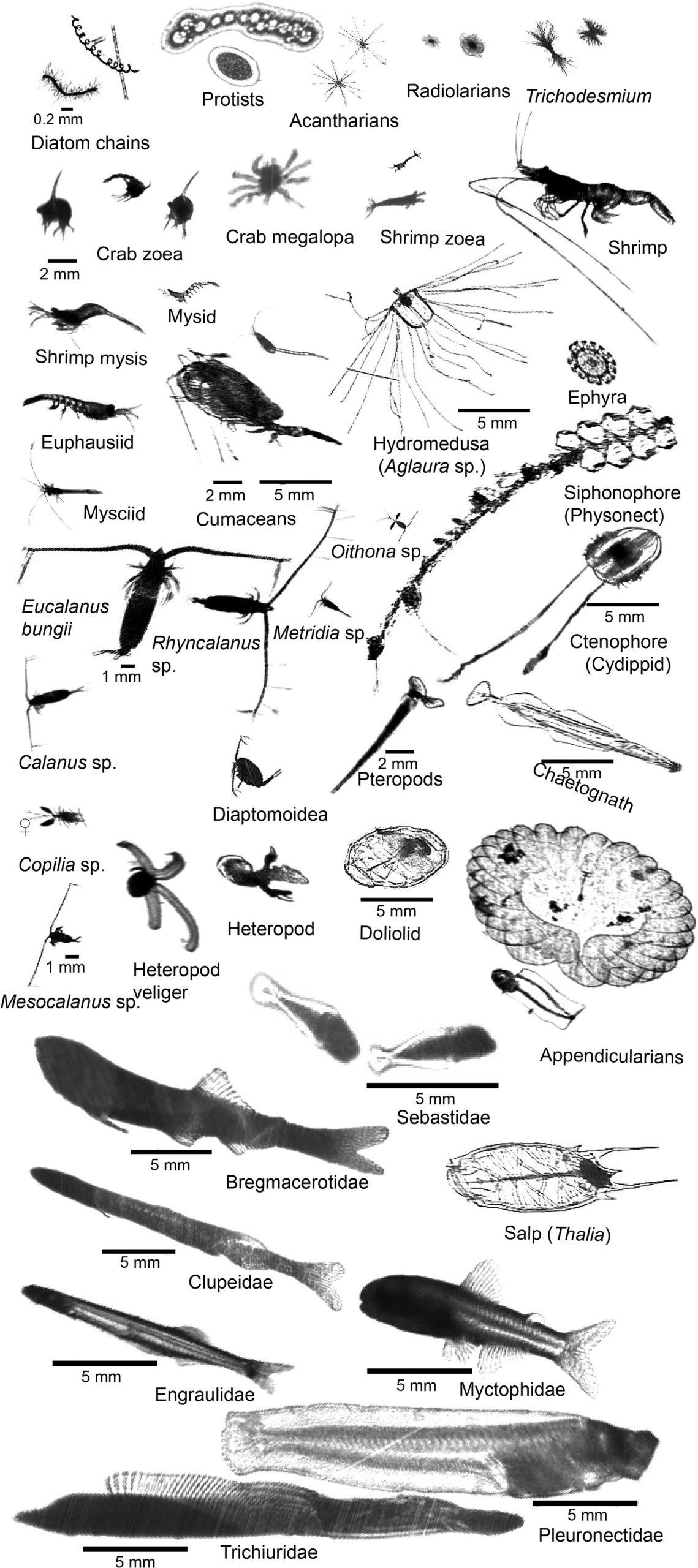
ISIIS-DPI images of key species in the Northern California Current including primary producers, protists, crustaceans, cnidarians, ctenophores, echinoderms, heteropods, pteropods, chaetognaths, pelagic tunicates, and larval fishes.
A team of scientists including Robert Cowen and Moritz Schmid of Oregon State University as well as Charles Cousin and Cedric Guigand, engineers from Bellamare LLC, thought up a better way to measure plankton populations. They developed the In-situ Ichthyoplankton Imaging System-3 (ISIIS-3), a kind of underwater sled they can fly through the ocean and which takes pictures of the microscopic life at different depths, with centimeter spatial resolution. But the pictures alone aren’t enough — identifying and counting the plankton in each image would take expert observers way too much time to catalog. Instead, they turned to artificial intelligence running on PSC’s NSF-funded, ACCESS-program-allocated Bridges-2 supercomputer and its AI-friendly CPU and GPU nodes.
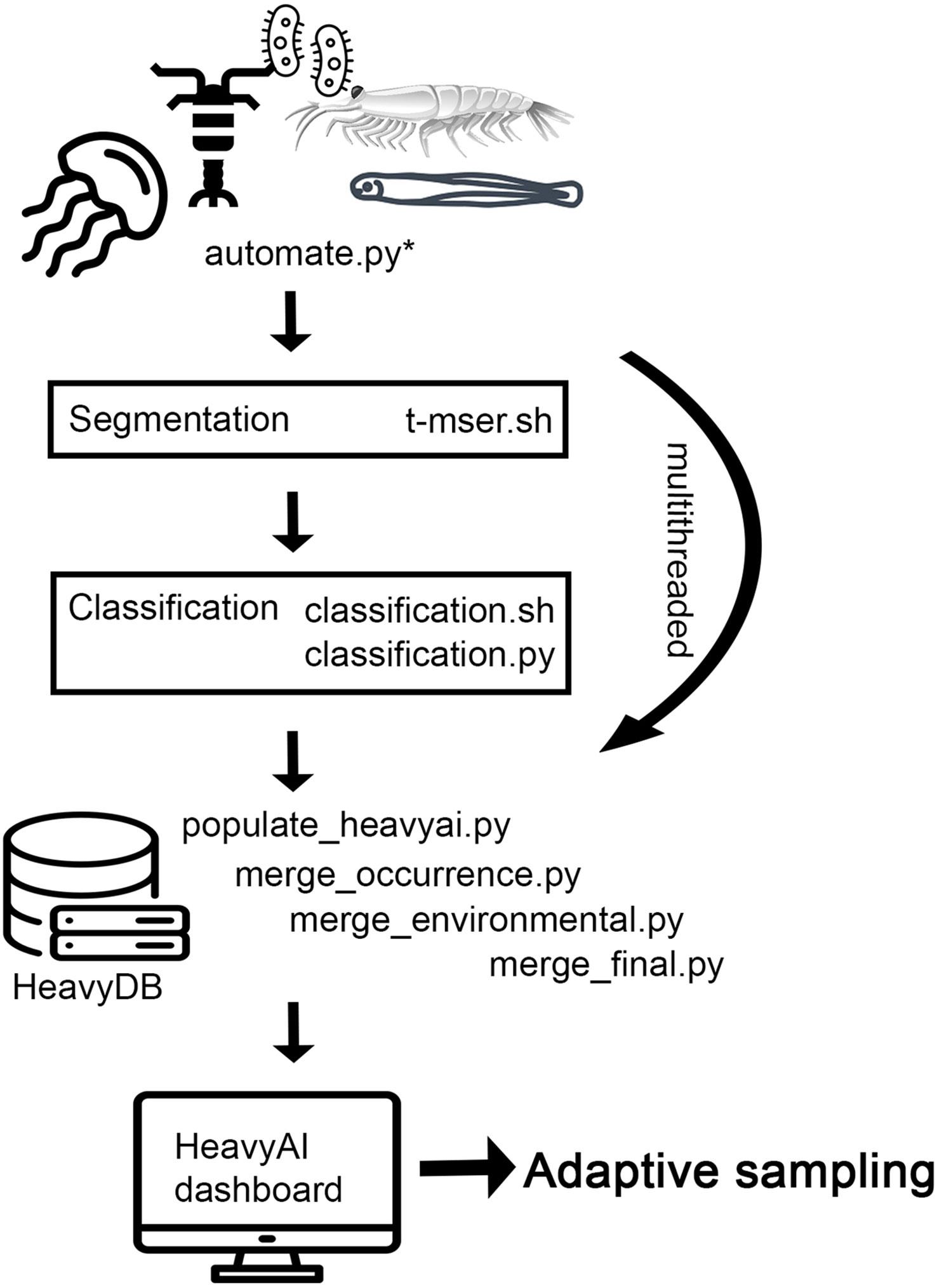
Pipeline schematic depicting the imagery data processing pipeline deployed at sea.
HOW PSC HELPED
The first challenge the team encountered was the volume of data that ISIIS-3 generates. In a five- to 10-day ocean cruise, the system would produce something between 40 and 80 terabytes of data. That’s enough to fill the hard drives of 40 or more high-end laptops. Moving such “Big Data” in and out of its processors without big time lags is a particular strength of Bridges-2. The work would have to take place in two major steps.
First, they would need to segment the images. That means pulling out “things” from the image that are not just background water. Accurately identifying a chunk of pixels in an image as something isn’t easy. It required the raw number-crunching ability of Bridges-2’s CPU (central processing unit) nodes.
Second, the team would need to classify what each of the segmented items is. This is where the AI comes in. By giving the AI more than 90,000 images in which the microbes had been identified and labeled by human experts, the scientists allowed the software to train itself. Then the scientists could test it against images without the labels, so that it corrects its mistakes and learns how to identify the plankton accurately. Bridges-2’s late-model GPU (graphics processing unit) nodes, a technology originally developed to give sharper images in video games, are particularly good at this kind of visual identification task. The scientists say that PSC’s Roberto Gomez and Sergiu Sanielevici were invaluable in making the system work.
“We started with PSC; that’s where the resources are that we need to get through this data in a timely manner, to close the time gap between collecting data and being able to work with the classified data … When we talk about PSC and going out with the edge server, while that’s not PSC infrastructure, [Bridges-2] is such a great resource that … the need to classify big data won’t go away.” — Moritz Schmid, Oregon State University
The trained AI proved to be reliable in identifying plankton in the ISIIS-3 images. That led to an even more ambitious goal. Could they export their software to a powerful computer that was nonetheless compact enough to run onboard a ship? ISIIS-3 and the AI had enabled them to carry out deep analyses of plankton populations using Bridges-2 once they came back to port. But at that point it would be too late to, say, turn the ship around to get another look at a particularly interesting location and depth. Onboard edge computation could solve that problem.
The team tested their AI at sea, showing that it could differentiate about 170 different plankton categories accurately and quickly enough to allow for “adaptive sampling,” enabling them to redirect the cruise where the data led them. First author Moritz Schmid of Oregon State and his colleagues reported these results in Frontiers in Marine Science in June 2023.
The scientists aren’t done with Bridges-2 or PSC yet. While adaptive sampling is a big leap forward in their ability to collect data, they still need to do hard-core processing when they get back ashore. One cruise easily leads to more than 2 billion plankton images. In addition, using PSC as a “one stop shop” will make it easier for them to share their tools with their colleagues at other institutions. One of the future goals of the project will be for graduate student Elena Conser to study how areas of low oxygen off the Oregon Coast affect which plankton survive and at what levels their populations persist.
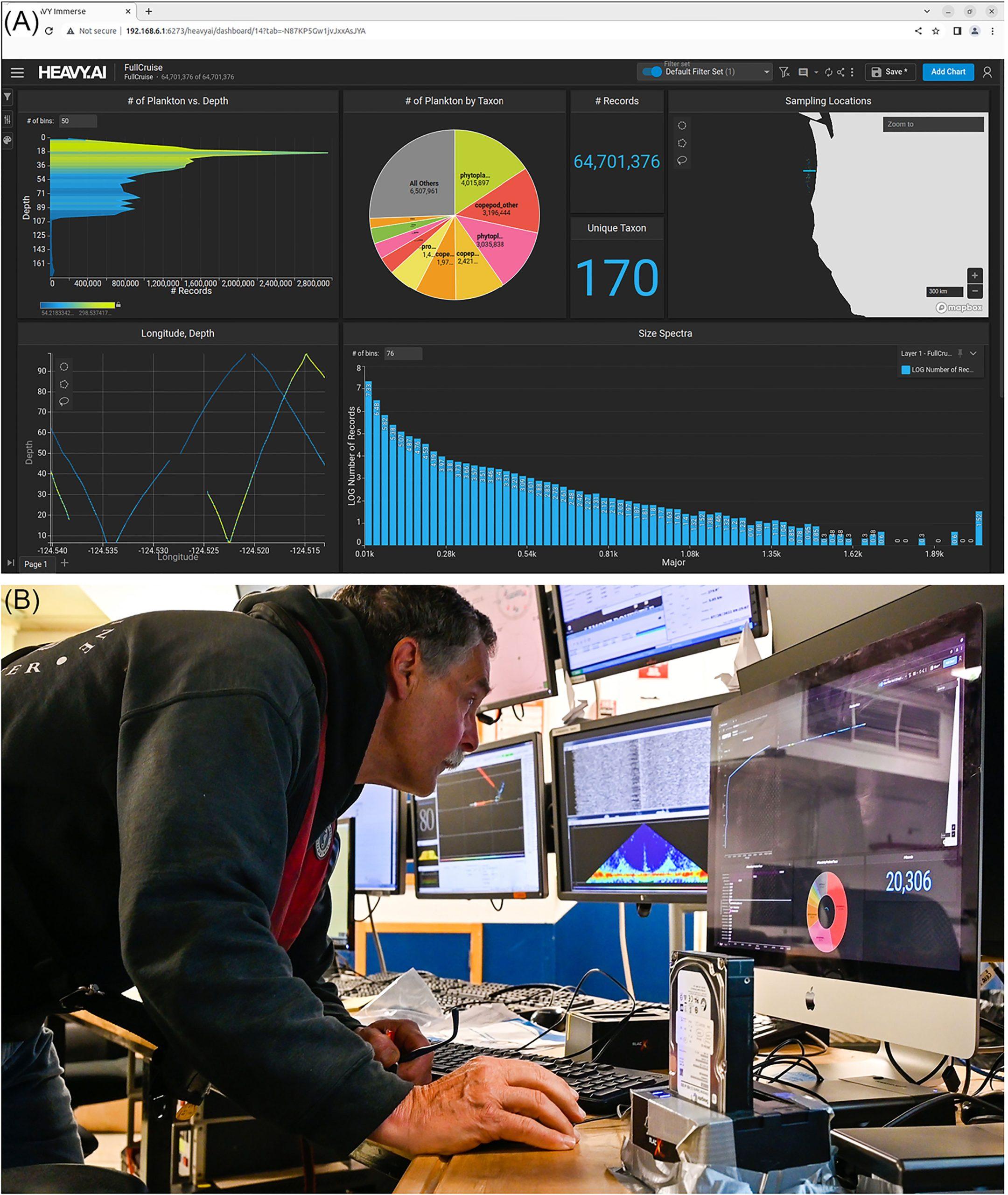
(A) The HeavyAI dashboard displayed on the adaptive sampling display used to analyze the ISIIS-DPI images; (B) This setup lends itself to near real-time data exploration and adaptive sampling.
