Nonidentical Twins
Anton 2 Simulations Show Curious Difference in Action of Paired Neurotransmitters
The NMDA receptor protein underlies memory formation. It’s also an important target of drugs for conditions as different as depression, Alzheimer’s disease and tinnitus. Scientists might be able to design better medications for these conditions if they understood better how the receptor works. Using Anton 2 at PSC, Johns Hopkins scientists simulated the receptor and the neurotransmitter chemicals that activate it. They discovered a curious difference in how the different neurotransmitters work that may be biologically important and offers new targets for drug development.
Why It’s Important
What was the first telephone number you learned? What’s the capital of France? What did you have for dinner yesterday? What do you need to pick up on your way home from work tomorrow?
Being able to answer these questions—forming and retrieving memories—depends on the nerve cells in your brain strengthening some of their interconnections and weakening others. This phenomenon underlies our ability to learn and think.
A central part of that ability is a protein called the NMDA receptor. This barrel-shaped protein sits in the outer membranes of brain nerve cells at the points where they connect and communicate with each other. When the neurotransmitters glutamate and glycine—with glycine sometimes replaced by D-serine—float across the space between two nerve cells, called the synapse, they together cause the NMDA receptors in the second cell to open a channel in their centers. The open channel lets electrically charged ions cross the membrane between the outside and the interior of the nerve cell. The flow of ions in turn activates the nerve cell, and can make it either more or less likely to be activated in the future. In other words, strengthening or weakening the cells’ connection.
“In vertebrates, there are three main functional glutamate receptors [in nerve cells]. The NMDA receptor … does not operate the way the others do. It’s unique in that you need both glutamate and either glycine or D-serine to activate it. We wanted to understand in terms of structure and function how are NMDA receptors different from the others?”—Albert Y. Lau, Johns Hopkins University School of Medicine
The NMDA receptor is important for memory formation. But it also plays a central role in a number of medical conditions. Doctors use drugs designed to affect the NMDA receptor for conditions as different as depression, Alzheimer’s disease, and tinnitus (ringing of the ears). Scientists also use it as a tool to create a condition similar to schizophrenia in rodents, so they can test anti-schizophrenia drugs.
Despite its importance, scientists didn’t know much about how glutamate and glycine bind to the NMDA receptor. Why are two different neurotransmitters necessary? How can D-serine sometimes substitute for glycine? Answering these questions—seeing more clearly how the neurotransmitter molecules interact with the receptor—could lead to better-designed medicines with fewer side effects. Alvin Yu, now a postdoctoral fellow at The University of Chicago, worked with advisor Albert Y. Lau at Johns Hopkins University (JHU) School of Medicine to try to find out more about these interactions by simulating the molecules in computers.
How PSC Helped
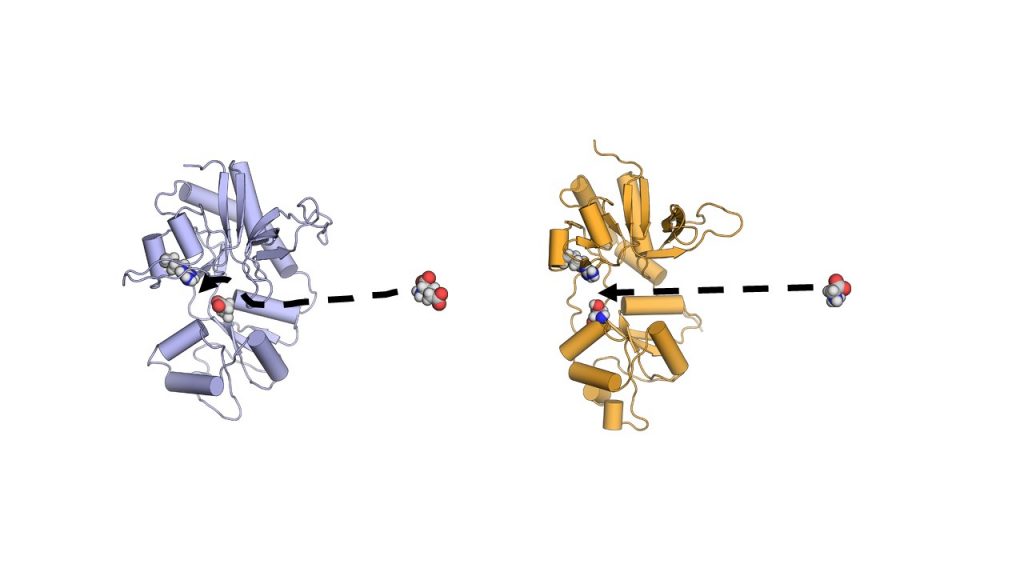
As part of his PhD work, Yu simulated how glutamate, a relatively large amino acid, and glycine, the smallest one, approach and stick to the NMDA receptor. He employed the MARCC supercomputer at JHU and used the string method to examine the motions and interactions of the molecules. He found something surprising. In the simulations, glutamate and glycine acted completely differently. Glycine just floated directly into the receptor’s binding site. Glutamate, on the other hand, had complicated interactions with parts of the protein just outside of the binding site. Like arms reaching out to grab the neurotransmitter, the “sticky sidechains” of the protein guided the glutamate molecule on its way in. The latter was a phenomenon that the scientists had seen in earlier work on another glutamate-activated nerve cell protein, the AMPA receptor.
But the JHU team wasn’t completely satisfied with the results. The string method depends on a lot of assumptions about how the molecules will act. They wanted to get an “unbiased” look at what the molecules would do without all the constraints. Their goal was to allow the virtual molecules to associate with each other using only the laws of physics that govern chemical interactions, without the assumptions of the string method. The MARCC supercomputer wasn’t powerful enough for these simulations.
To get the power they needed, the scientists turned to Anton 2 at PSC, which is specialized for such “molecular dynamics” simulations. Anton 2 is hosted at PSC thanks to manufacturer D.E. Shaw Research and operational funding by the National Institutes of Health.
“At Johns Hopkins we have a high performance computing resource called MARCC. It’s a terrific resource … however, for the time scales that we wanted to access for these particular studies, Anton was really the best resource we had available. Our initial studies on isolated ligand-binding domains involved maybe 40,000 atoms. Now on Anton 2, we’re running simulations of the entire receptor embedded in the membrane—monster systems with around 700,000 atoms.”—Albert Y. Lau, Johns Hopkins University School of Medicine
Using Anton 2, the team was able to carry out more than 50 microseconds of simulations in about a week of real computing time. The same computation would have taken 10 years on a general-purpose computer cluster. Even better, the unbiased view that Anton 2 gave them of the NMDA receptor and its neurotransmitters agreed with the enhanced sampling simulations on MARCC. Glycine moved straight to its destination. Glutamate followed a less direct route, with the protein’s sticky side chains guiding it in. The scientists could now be more confident that the difference between the two neurotransmitters’ routes was real, rather than a product of their assumptions. The team reported their results in the journals Neuron and Structure.
The JHU researchers aren’t yet sure why the pathways the two neurotransmitters take differ. Since glycine is smaller than glutamate, one possibility is that it just has an easier time sliding in and doesn’t need the help. The helping hands that glutamate experiences might be a way for nature to speed up the important communication between nerve cells. Glycine may just be too small to form these interactions. Another question they’d like to answer is whether drug-like molecules bind like glutamate and glycine, or by some other mechanism. They plan to attack these problems with Anton 2.