Training & Education
PSC offers training in HPC, Big Data, bioinformatics, biomolecular simulation, and more. Workshops address topics ranging from code optimization and parallel programming to specific scientific interests.
PSC workshops and STEM education resources incorporate lectures and extensive hands-on sessions, along with online teaching tools, online courses, and workshop webcasts, among other approaches. Programming exercises are carefully designed to reinforce concepts and techniques taught in class.
Instructors have strong scientific, technical, and design backgrounds and are available for consultation, including help with participants’ own coding needs and curriculum building.
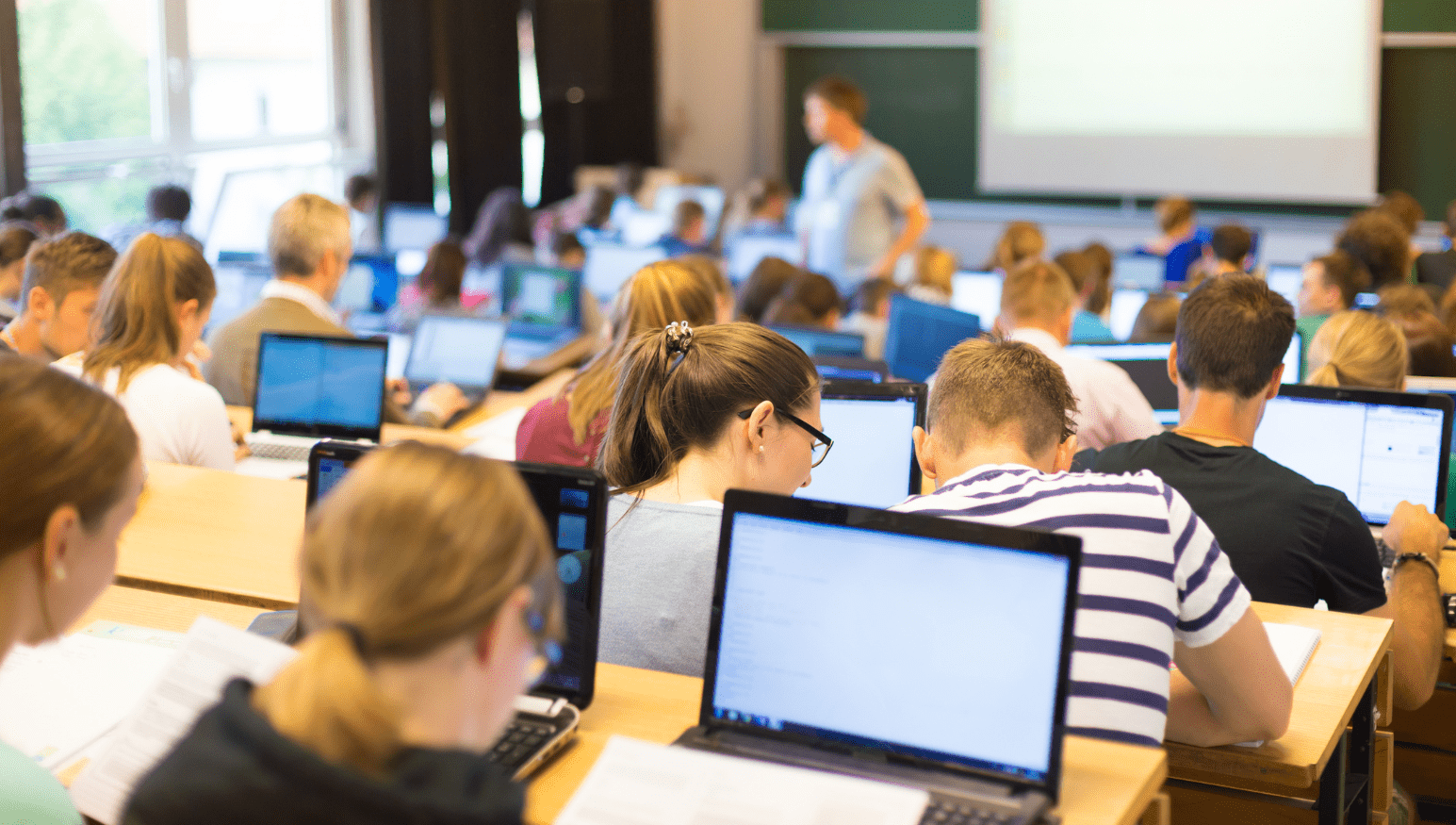
High Performance Computing workshop series
This renowned series educates hundreds of students monthly on a variety of HPC topics, including Big Data, and GPU and parallel programming. Workshops are broadcast live to dozens of satellite sites, and have reached more than 10,000 participants in five years.
PSC Learning Lab
The PSC Learning Lab provides a curated collection of educational resources designed to elevate your skills in high-performance computing, data science, and research technologies.
Learn at your own pace and grow with courses designed by experts.
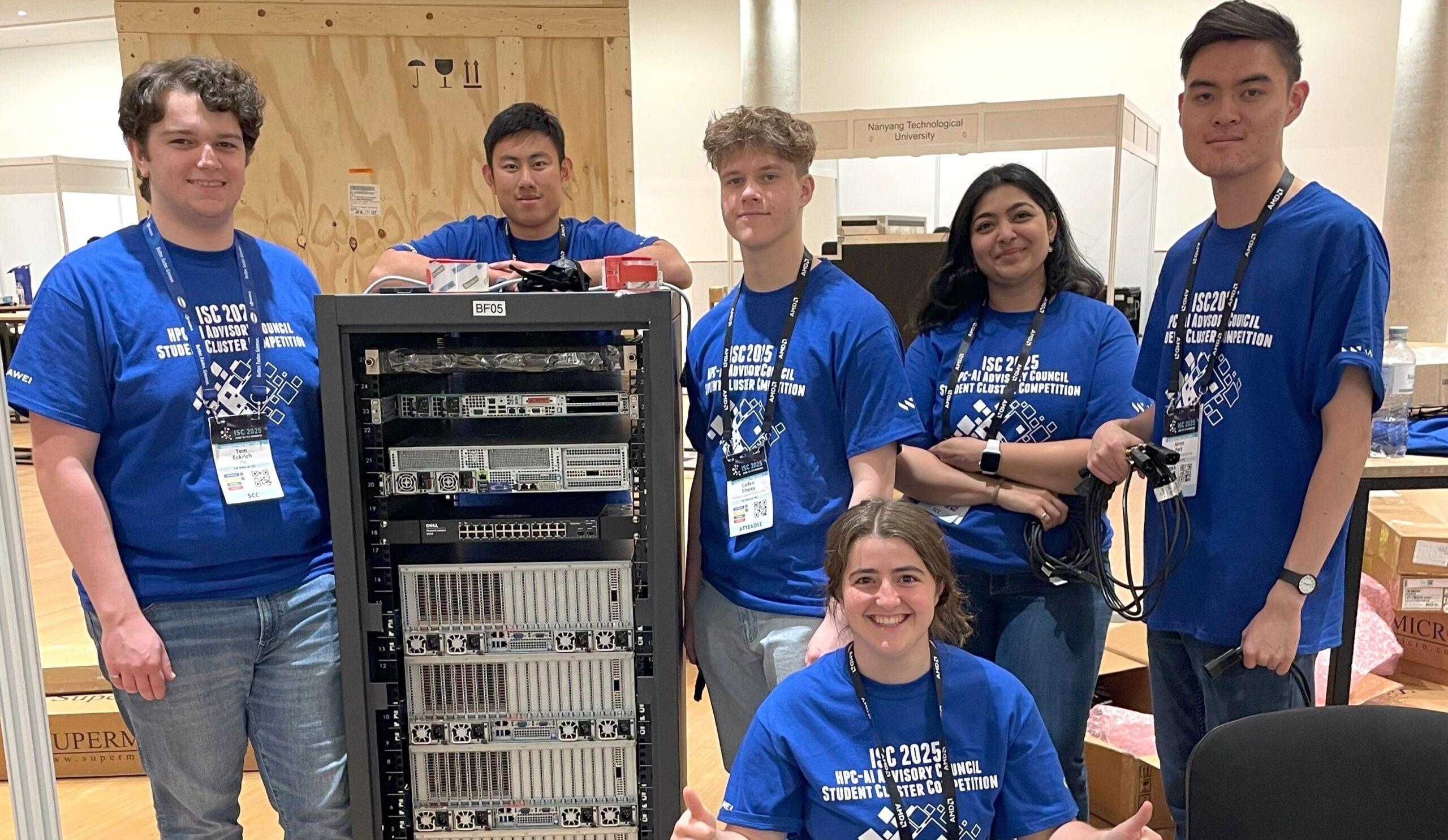
Internship program
PSC’s Internship Program supports workforce development in the state of Pennsylvania. Through this program, undergraduate students gain valuable experience doing research and working on development projects in areas such as data analytics, computational science, and bioinformatics with some of the world’s leading experts in their fields.
Check our careers page for open internship positions.
Bridges-2 webinar series
The Bridges-2 webinar series presents topics of interest to enable the Bridges-2 user community to optimize and accelerate their research.
Neocortex training
Training on the Neocortex system presents a system overview of this innovative platform for AI and ML research, and insights on how to best take advantage of its unique hardware, software, and capabilities.
Anton
Training on Anton, a special purpose supercomputer for biomolecular simulation, designed and constructed by D. E. Shaw Research, is conducted yearly for researchers who have received an Anton allocation.
Cybertraining: ByteBoost
Byte Boost offers experiential learning opportunities on modern, cutting-edge, specialized computing technologies. This initiative will empower researchers to make the best computing choices in the ever-changing landscape of computational technology by focusing on supporting researchers to use the technologies of three NSF supported computing testbeds – Ookami at Stony Brook University; Neocortex at PSC; and ACES at Texas A&M University.
CI Pathways
CI Pathways is a cohort-based training program offering a guided approach to use Cyberinfrastructure to enhance research.
Learn alongside peers and receive expert guidance from seasoned mentors. Participants will have opportunities to build relationships lasting beyond the CI Pathways program.
ADAPT PA: College-level curriculum development
In the ADAPT PA program, PSC partners with community colleges and PASSHE schools across the state of Pennsylvania to enhance curricula, provide learning resources, and collaborate on new programs.

STEM programs
PSC STEM programs and educator resources provide online content for K-12 teachers and students. Many also offer workshops (with Act 48 hours) on how to effectively use the materials. PSC features pre-college programs, resource libraries, and bioinformatics activities across multiple disciplines. For more information on incorporating these materials into a workshop or professional learning experience, contact eotinfo@psc.edu.
Bioinformatics
We offer summer workshops on various bioinformatics topics.